DROMA
Drug Response Omics association MAp (DROMA, 卓玛)
DROMA is a comprehensive database and analysis tool that integrates the largest published studies investigating cancer response to chemical compounds and the associations between drug sensitivity and multi-omics data (mRNA, CNV, protein, mutation, etc.) across various cancer models including PDC (Patient-Derived Cells), PDO (Patient-Derived Organoids), and PDX, human data are under development.
Main components
- Projects about data:
DROMA_DB: mugpeng/DROMA_DB: Sqlite db and DromaSet obj for Drug Response Omics association MAp. (DROMA)
DROMA_Set: mugpeng/DROMA_Set: DROMASet is a comprehensive R package for managing and analyzing drug response and omics data across multiple projects. It provides a robust framework for handling complex multi-omics datasets with integrated drug sensitivity information, enabling seamless cross-project comparisons and analyses.
- Projects about analysis:
DROMA_R: mugpeng/DROMA_R: R package for DROMA.
DROMA_Web: mugpeng/DROMA_Web: Shiny website for Drug Response Omics association MAp. (DROMA)
- AI and AI agent:
DROMA_MCP: [mugpeng/DROMA_MCP: DROMA MCP Server bridges the gap between AI assistants and cancer pharmacogenomics analysis by providing a natural language interface to the DROMA.R and DROMA.Set packages.](https://github.com/mugpeng/DROMA_MCP)
- Under development:
DROMA_MCP, DROMA_AI, DROMA_Augur, DROMA_py
Statistics Info
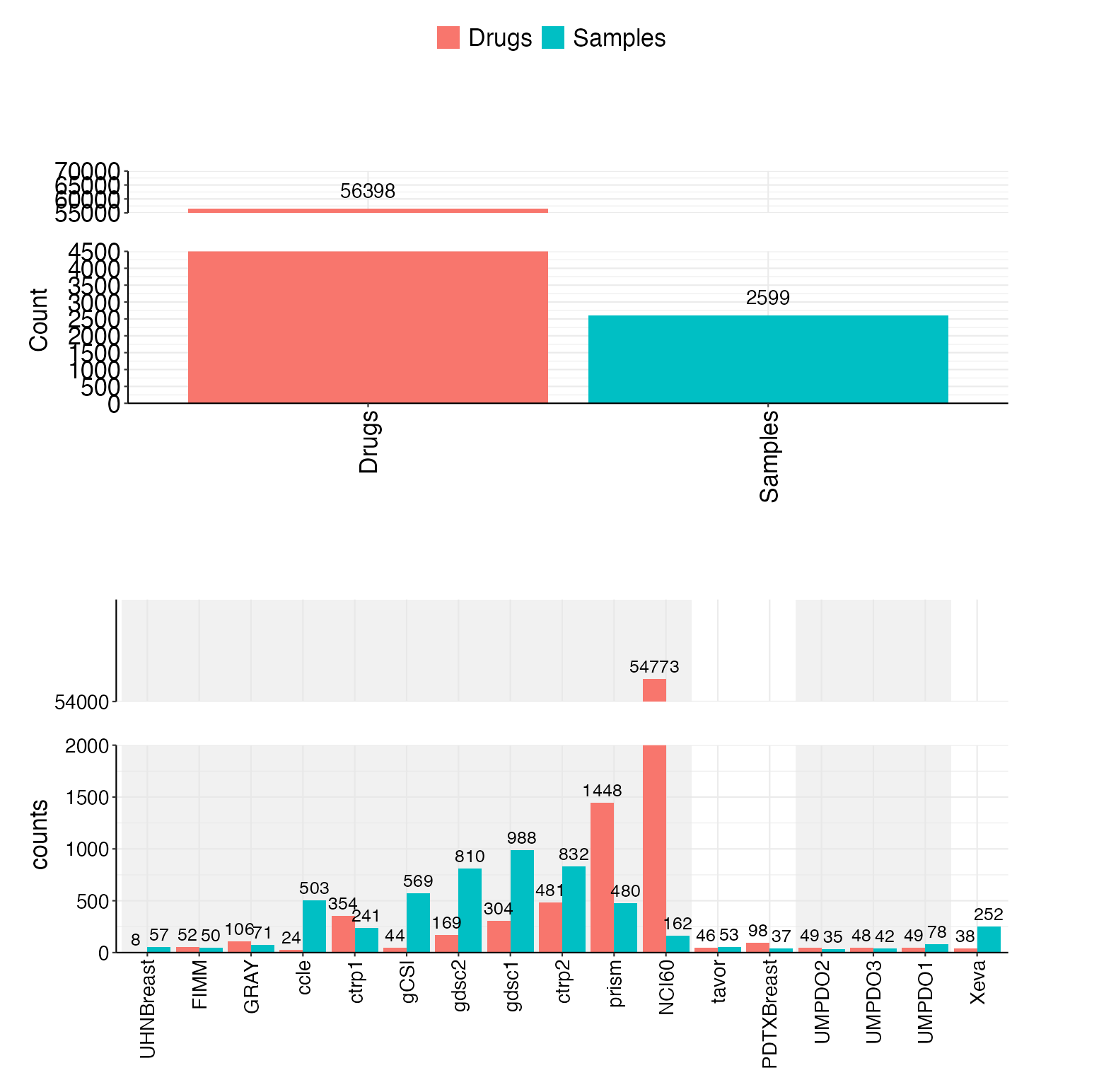
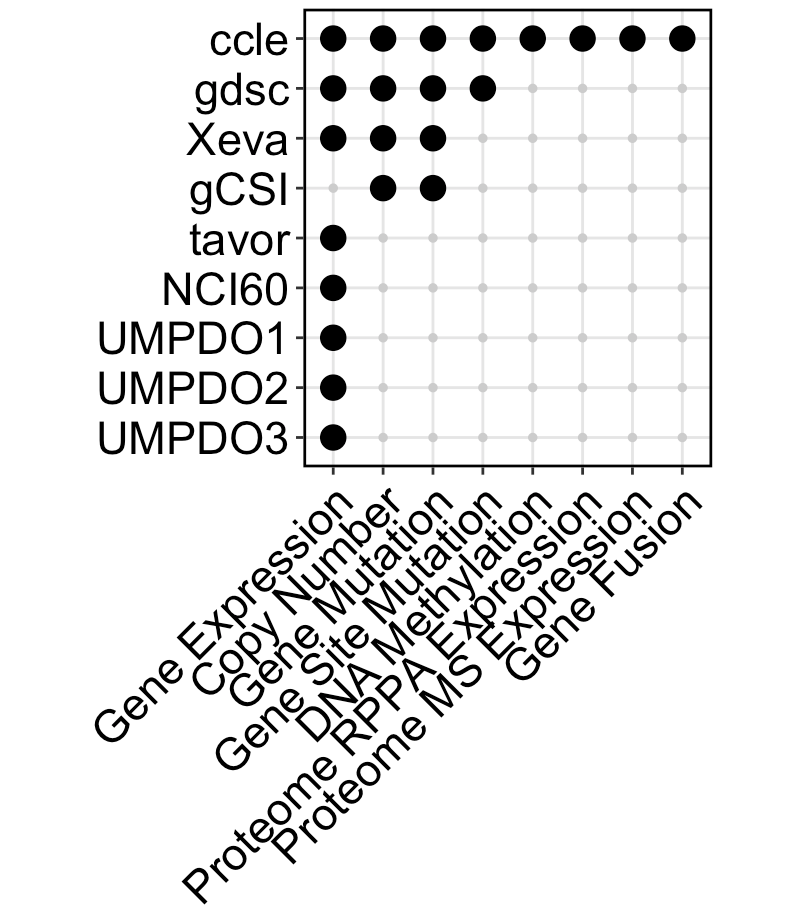
Include 17 datasets now(11 cell line, 2 PDC, 3 PDO, 1 PDX), totally 2599 unique samples and 56398 unique drugs.
Citation
If you use DROMA or any subprojects(DB, R, web…) in your research, please cite:
Li, S., Peng, Y., Chen, M. et al. Facilitating integrative and personalized oncology omics analysis with UCSCXenaShiny. Commun Biol 7, 1200 (2024). https://doi.org/10.1038/s42003-024-06891-2
Milestone
0526 v0.4 andata
坂本龍一:【氛围感|andata (Electric Youth Remix)】 https://www.bilibili.com/video/BV1v7kAY7Eoi/?share_source=copy_web&vd_source=bd40fa636d7a7f4f2a51596fed85e2b0
0513 v0.3
Refactor App.R and Modules: Update version to 0.3.
- add new datasets, now have in vivo data now(Xeva PDX dataset), totally include 17 datasets now(11 cell line, 2 PDC, 3 PDO, 1 PDX)

- add new module, and enhance drug feature analysis.
- Remove deprecated Rmd files and adjust data loading paths.
- Update BatchFeature and DrugOmicPair modules for improved functionality and user experience.
- Functionize all functions in modules and put them under
Package_Function/preparing for making DROMA R package. - fix some bugs.
0319
I will attend UM PhD seminar, welcome!
0318 v0.2
The stable version v0.2 is online now in UM network: http://fscpo.fhs.um.edu.mo/DROMA_DB/ Btw, please be patient, it may take 10-20s when first time start it. :)
This update includes:
- Add denglab three PDO data(deng1(breast), deng2(colon), deng3(nasopharynx)) with paired RNAseq, IC50 and annotation data;
- Harmonize cell line drug and cell names also the annotation data;
- Add choice to filter data for a specific tumor type(Breast, lung…) or data type(cell line, PDO), and a global setting page to set data. Now all continous data is transformed to by z-score in default, which allow to add a merged result for visualization ;
- Add several figures and some visualization-related improvement.
DROMA MCP Server
Project Details
- mugpeng/DROMA
- Mozilla Public License 2.0
- Last Updated: 6/4/2025
Recomended MCP Servers
Model Context Protocol server for GitHub API integration
Model Context Protocol (MCP) server for BatchData.io property and address APIs - Real estate data integration for Claude...
This repository hosts the “Weapon Recoil Generator” MCP Server deployed on Smithery.
A MCP server for Google Analytics Data API
QMT-MCP-Server
A Model Context Protocol (MCP) server that enables AI agents to interact with the Tally API, providing access...
Integrate librosa, whisper with LLMs to analyze music audio.